|
Dhruva Abhijit Rajwade
I am working with Prof. Marianna Rapsomaniki's AI/ML For
Biomedicine
group
in CHUV, Lausanne as a RA, working on drug
perturbations, causality and and generative modeling for
biology. I graduated in May'25 as an Integrated Master's
student at
IIT Kharagpur in
India, where I Majored in Biotechnology and Biochemical
Engineering and worked in Computational Biology,
Computer Vision and AI4Science.
In my undergrad journey, I have worked with Prof.
Koel Chaudhary
and Prof.
Soumya De
at IIT Kharagpur. In the summer of 2023, I got an
amazing opportunity to work with Prof.
Brian Ingalls
at the University Of Waterloo. I worked at Caltech in
the summer of '24 with Dr.
Shengchao Liu
on understanding Protein-DNA interactions using
Cross-Attention and language modelling. For my Master's
thesis, I extended my work at Caltech by using discrete
diffusion for controllable generation of DNA-binding
protein sequences, with a focus on discovering novel de
novo DBPs. Some of my designs expressed well in the lab,
and this is an ongoing project in IIT Kharagpur at Prof.
Riddhiman Dhar's
group.
I am a recipient of the
Caltech SURF
fellowship (2024), the
MITACS Globalink
Research Internship (2023), and have also been selected
for the
EPFL E3 scholarship
(2024) and the
ThinkSwiss
Research scholarship (2023).
Email
/
Scholar
/
Twitter
/
Github
|


|
Research
I'm primarily interested in Deep Learning for Protein
Design, Gene regulation and Biological systems,
Mathematical modelling of biological networks and
dynamics, and AI for science. Most of my work has
involved applying robust learning methods to Biological
problems in an interpretable manner. Apart from Biology,
I am keenly interested in and have also worked in Deep
Learning for Vision (specifically SSL, secure ML and
Generative modelling), as well as Causal Inference and
Graph Learning. More recently, I have been interested in
Fourier Neural Operators, Geometric Deep Learning,
State-space models and their applications in Biology.
|
|
[May 2025]
|
Some early work from my Master's thesis was accepted as a
poster at the
AI Bio X
conference at Sanger, Cambridgeshire. See you in the UK!
|
|
[Dec 2024]
|
Our
paper on
backdoor and adversarial attacks targetting SSL was
accepted at ICASSP 2025. See you at
Hyderabad!
|
|
[Nov 2024]
|
Our
work
on understanding Protein-DNA interactions using Protein
and Genomics Foundation models was accepted at the
MLSB, FM4Science and
AIDrugX workshops, NeurIPS 2024.
|
|
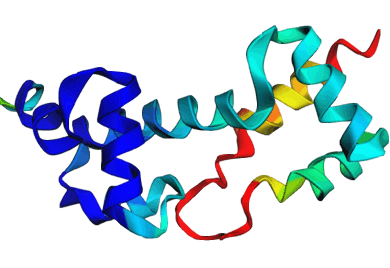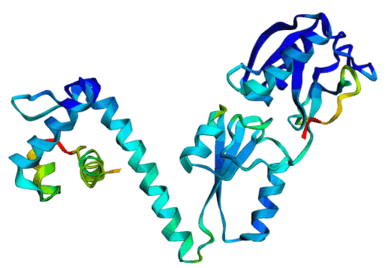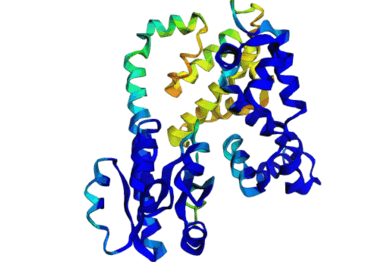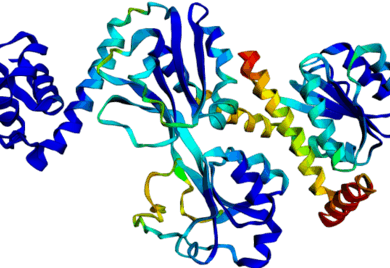
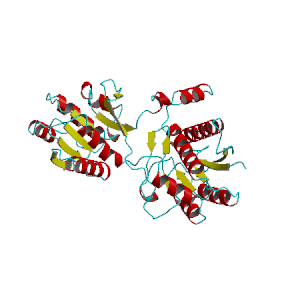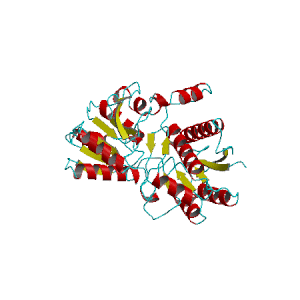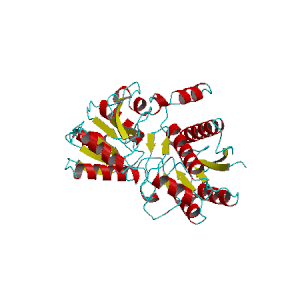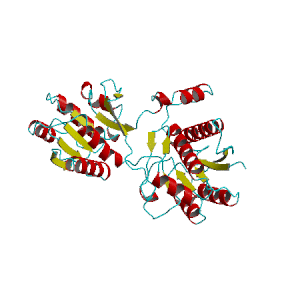
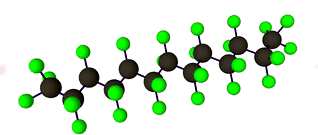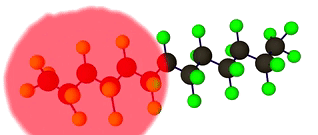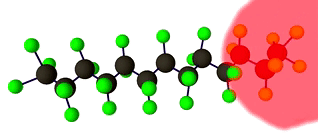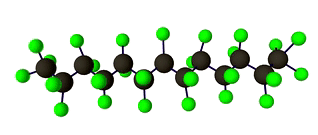
Discrete Diffusion For Tunable DNA-binding Protein
Design
[Slides]
Using Discrete Diffusion to learn the distribution of
DNA-binding protein sequences. Using our work on
Seq2Contact
to guide the diffusion process to generate proteins with
high affinity for specified DNA targets. Working on
optimizing CRISPR-Cas protein design, stitching together
different functional domains by inpainting a DNA-binding
domain in between, generating DNA sequences for binding
to a target protein by inverting the Seq2Contact model,
and engineering protein-DNA interactions. This project
is in collaboration with Prof.
Riddhiman Dhar
at IIT Kharagpur. Image shows ESMFold folded sequences
sampled by our trained discrete diffusion model, with as
less as 25% sequence similarity to the training set and
the presence of different DNA-binding domains (through
PFam scans). The protein sequences were not
cherry-picked, and the structures are coloured by
confidence (blue: High, red: Low).
|
|
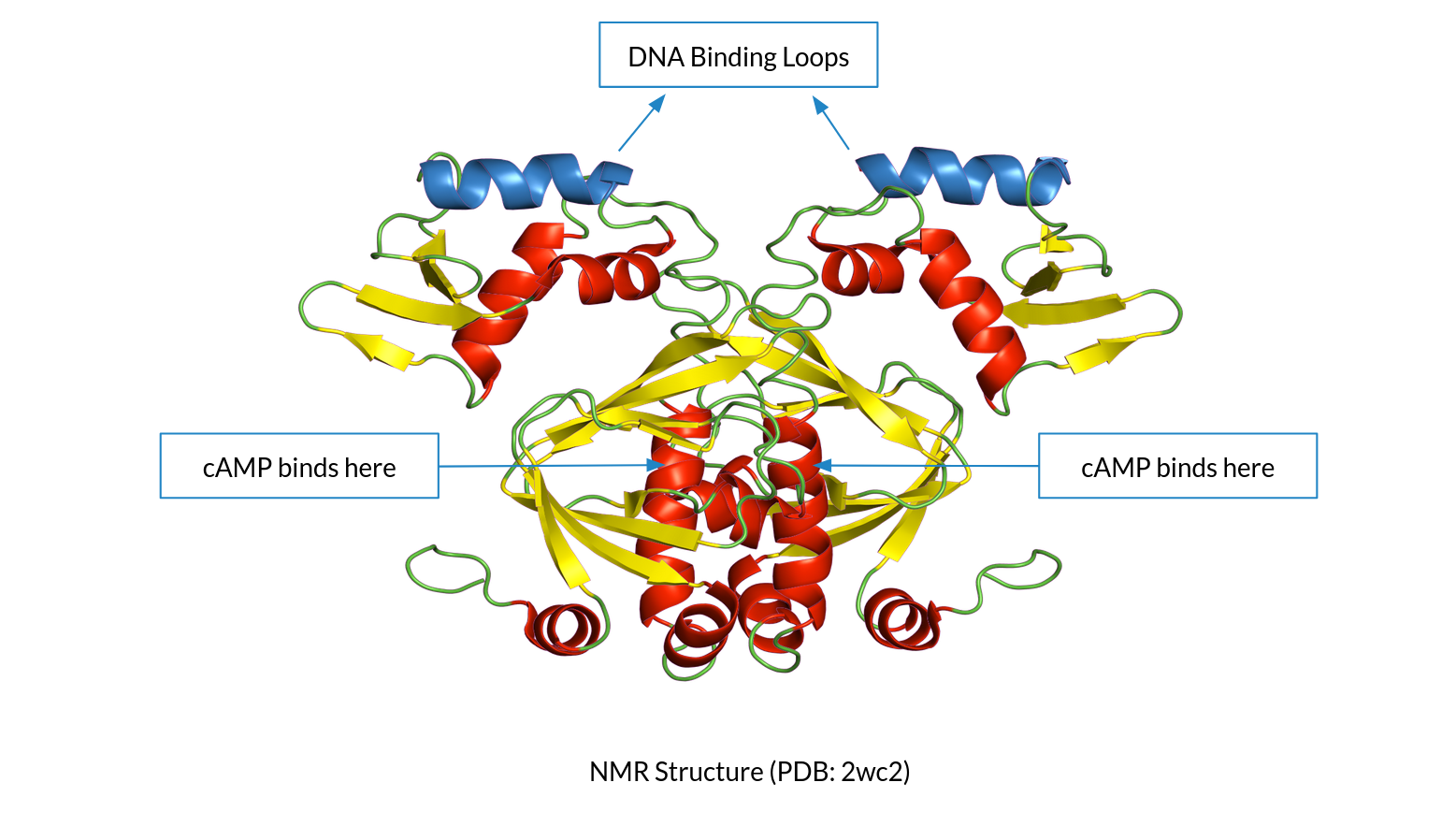
Finding Allosteric Networks in the CAP-cAMP system using
Deep Learning and Molecular Dynamics
[Slides]
Using MD simulations and trajectory analysis to infer
Allosteric networks in the CAP-cAMP system (where the
cAMP ligand binds to CAP protein, leading to allosteric
orientation changes and finally, transcription). Current
progress includes uncovering of three prospective
networks using
AlloReverse. Future plans include using State-space models to
learn the dynamics of protein-ligand interactions, and
finding a generalizable method for allosteric discovery.
This work is in collaboration with Prof.
Soumya De
at IIT Kharagpur.
|
[Note: Highlighted papers indicate first authorship; (*)
indicates equal contribution].
|
Understanding Protein-DNA Interactions by Paying
Attention to Protein and Genomics Foundation
Models
Dhruva Abhijit Rajwade, Erica Wang, Aryan
Satpathy, Alex Brace, Hongyu Guo, Arvind Ramanathan,
Shengchao Liu, Anima Anandkumar
NeurIPS 2024 Foundation Models For Science, AI for New
Drug Modalities, Machine Learning in Structural Biology
workshops, 2024
Paper
/
Code
Cross-Attention coupled with Protein and Genomics
Foundation models to understand Protein-DNA interactions
speeds up inference and achieves State-of-the-art
performance in predicting contacts in Protein-DNA
complexes (using purely sequence data for inference).
|
|
Towards Backdoor Mitigation and Adversarial
Robustness in SSL
Dhruva Abhijit Rajwade*,
Aryan Satpathy*, Nilaksh*
ICASSP 2025
Paper
/
Code
An intuitively elegant and simple defense strategy to
defend against standard SSL augmentation invariant
frequency based backdoor attacks. Taking a leaf out of
frequency domain attacks, we also use frequency domain
patching to increase model robustness in SSL.
|
|
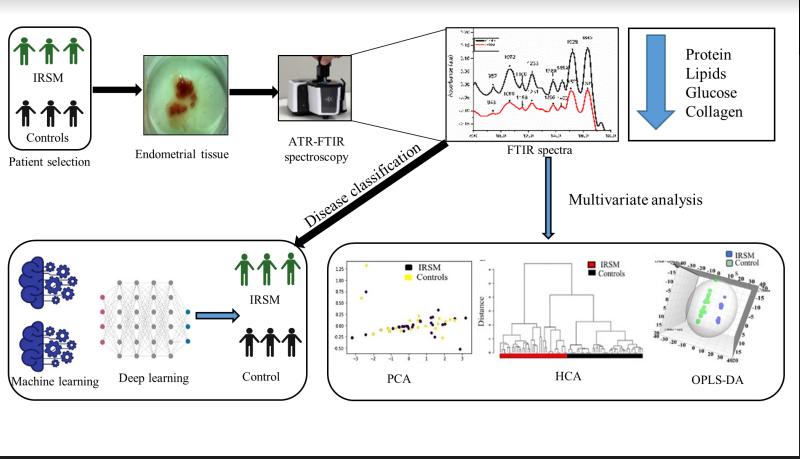
Attenuated Total Reflectance–Fourier Transform
Infrared (ATR-FTIR) Spectroscopy Combined With Deep
Learning for Classification of Idiopathic Recurrent
Spontaneous Miscarriage (IRSM)
Dadoma Sherpa, Dhruva Abhijit Rajwade,
Imon Mitra, Souvik Biswas, Sunita Sharma, Pratip
Chakraborty, Shovandeb Kalapahar, Ratna Chattopadhyay,
Koel Chaudhury
Analytical Letters (Journal), 2024
Paper
/
Code
An extension of our previous work (see below) on using
(ATR-FTIR)Spectroscopy and Deep Learning for prediction
of Idiopathic Recurrent Spontaneous Miscarriage (IRSM).
This work focuses on the classification of IRSM using
ATR-FTIR Spectroscopy, which is a non-invasive and
cost-effective technique and improves on our previous
work on using Raman Spectroscopy in a similar problem
setting.
|

|
Cells2Vec: Bridging the gap between experiments and
simulations using causal representation learning
Dhruva Abhijit Rajwade,
Atiyeh Ahmadi,
Brian Ingalls
NeurIPS 2023 Causal Representation Learning
Workshop, 2023
Paper /
Poster
/
Code
/
Slides
/
Talk
Learning meaningful representations of multi-cell
timeseries (Cellmodeller) simulations using causal representation learning.
Current work includes extending this to real-world data,
and for proxy-simulation generation for biological
experiments.
|
|
Prediction of Idiopathic Recurrent Spontaneous
Miscarriage using Machine Learning
[Best Paper Award]
Dadoma Sherpa, Dhruva Abhijit Rajwade,
Imon Mitra, Dhruba Dhar, Sunita Sharma, Pratip
Chakraborty,
Koel Chaudhury
IEEE International Conference on Computer, Electrical &
Communication Engineering, 2023
Paper
/
Code
Using Raman Spectroscopy and Machine Learning for
prediction of Idiopathic Recurrent Spontaneous
Miscarriage (IRSM). Improved this work using ATR-FTIR
Spectroscopy in a follow-up study, and currently working
on a multi-omics approach to the same problem.
|
|
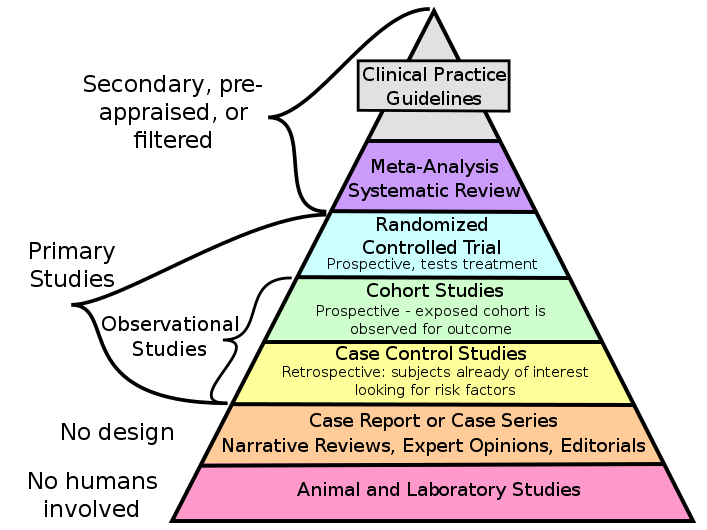
Risk factors associated with mortality in
hypersensitivity pneumonitis: a meta-analysis
Sanjukta Dasgupta, Anandita Bhattacharya,
Dhruva Abhijit Rajwade, Sushmita Roy
Chowdhury,
Koel Chaudhury
Expert Review of Respiratory Medicine (Journal),
2022
Paper
/
Code
Using different statistical tests and empirical analyses
to identify risk factors associated with mortality in
Hypersensitivity pneumonitis, a rare lung disease.
Checking for Publication bias and heterogeneity in the
data, and using meta-analysis to combine results from
different studies.
|
|
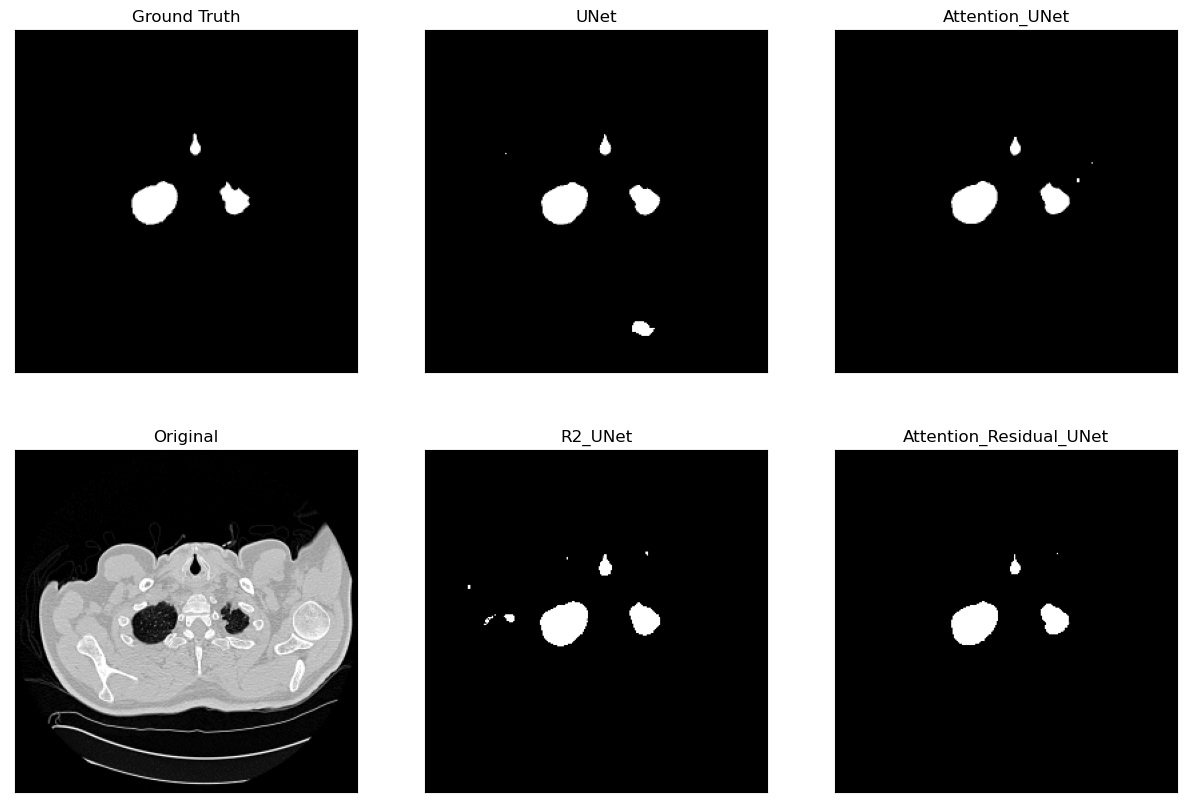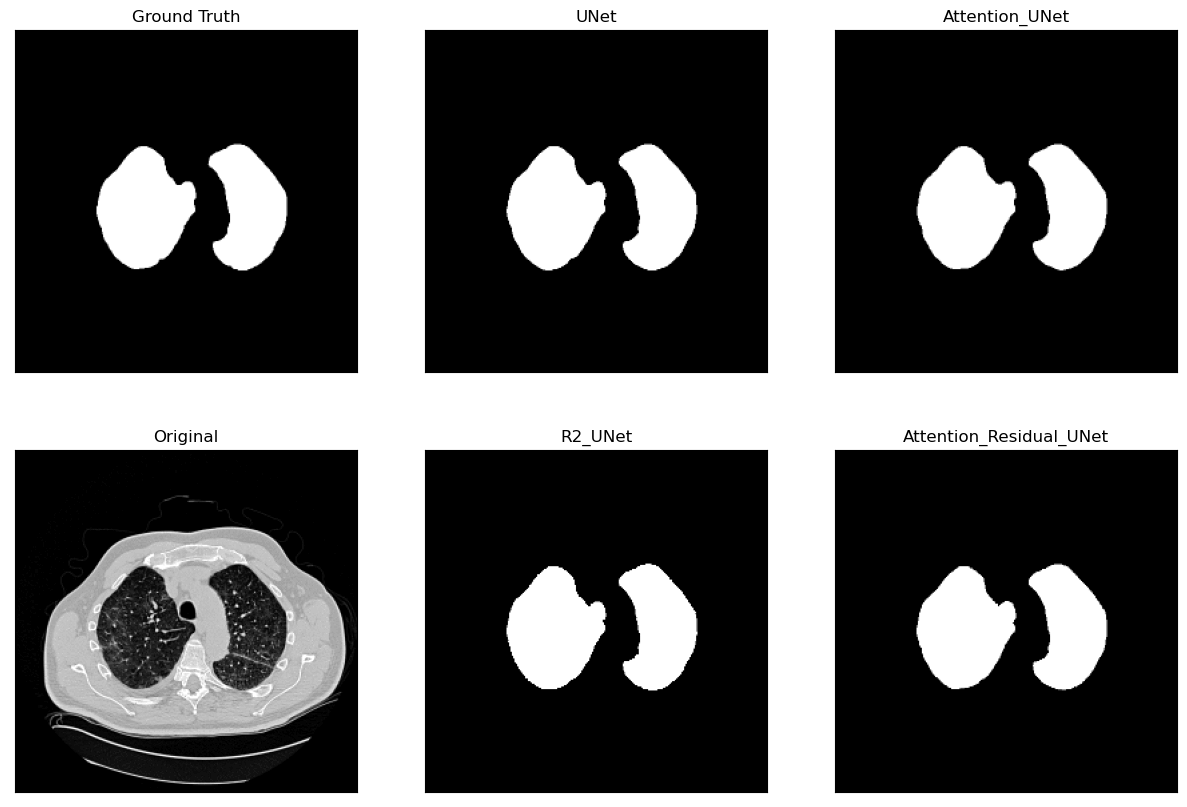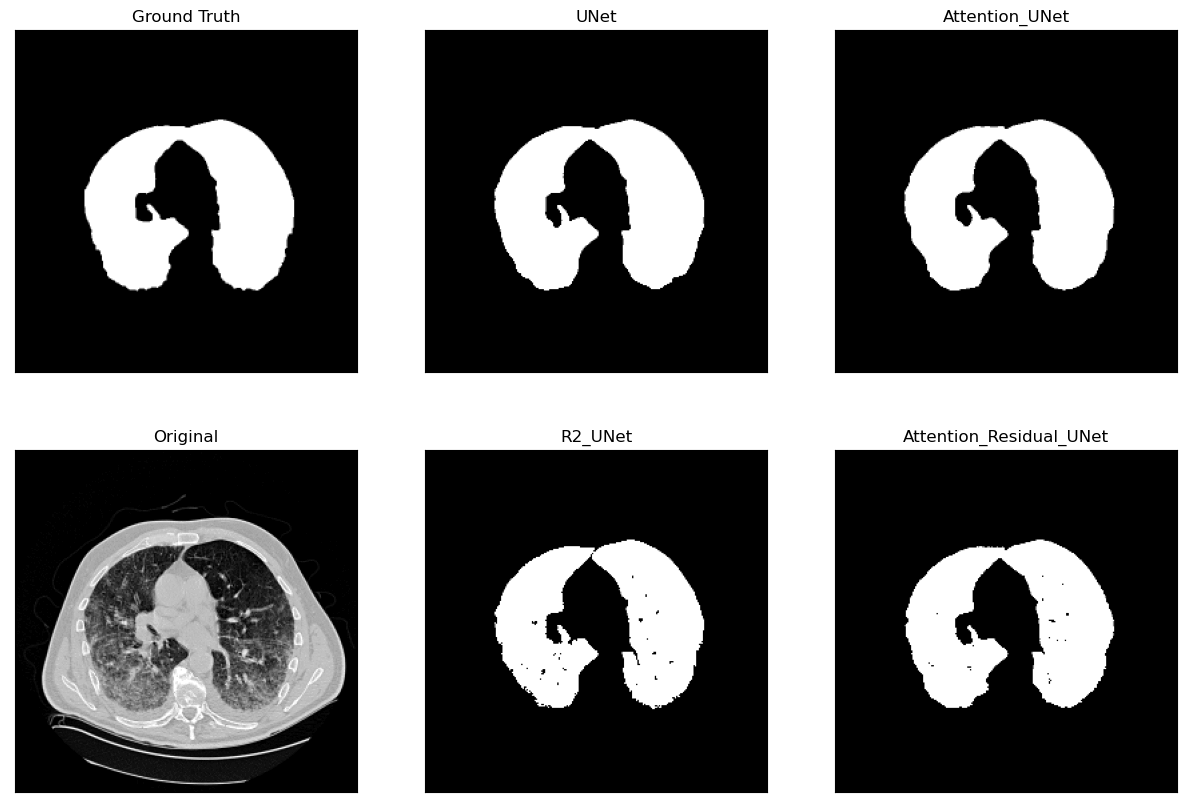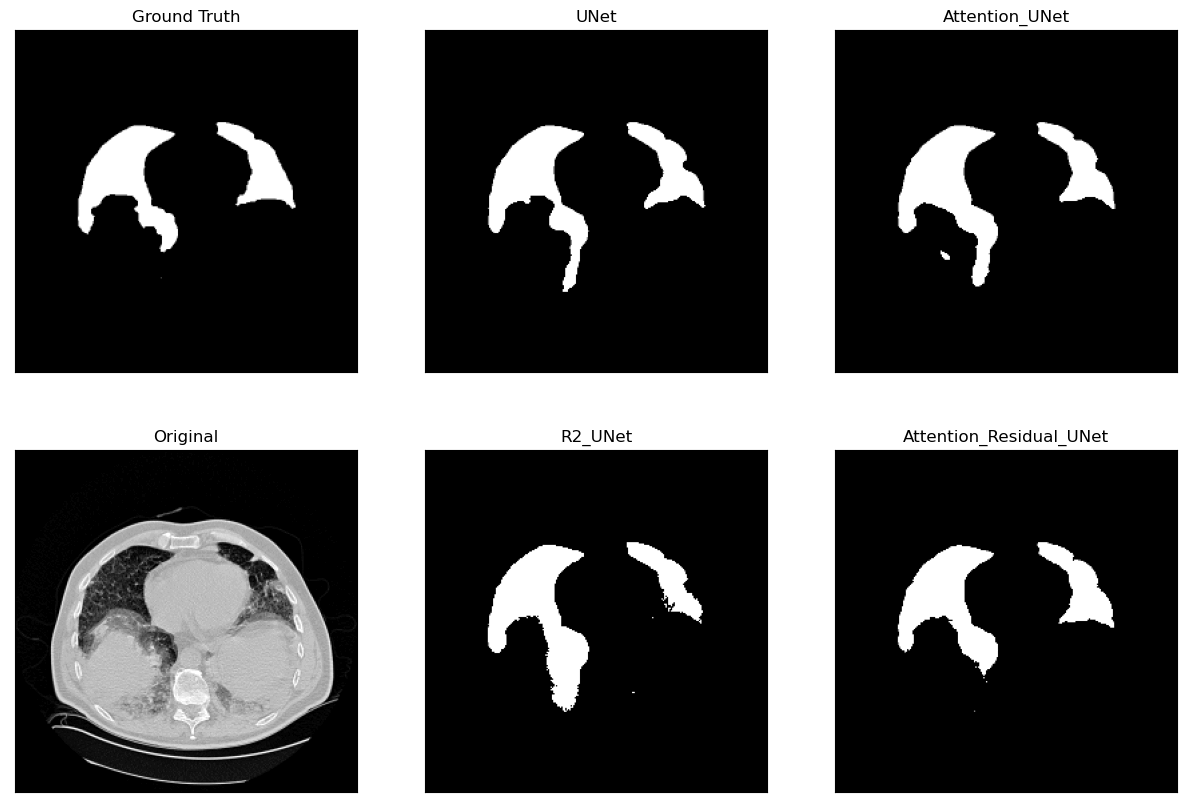
Lung Segmentation and Disease Classification using Deep
Learning
[Code]
Worked on using the
Geneva HRCT dataset
to segment lungs and classify diseases using Deep
Learning. The final model uses a U-Net architecture for
segmentation and a CNN for classification. The model was
trained on a subset of the dataset and tested on CT scan
images obtained externally through collaborations with
Hospitals.
|
|
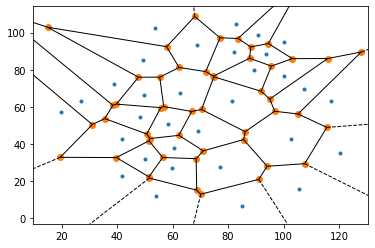
Functional Network Analysis of Calcium ion pathways in
beta-islets of the Pancreas
[Code]
Worked on using Calcium ion imaging time-series data to
extract functional networks in beta-islets of the
pancreas. Used Voronoi Delaunay triangulation (see
image) to extract the network, and used graph theory to
analyze the network. Code is incomplete and will be
updated soon.
|
This website's template is taken from
Jon Barron. Do not scrape the HTML from this
page itself, as it includes analytics tags that you do
not want on your own website — use the github code
instead. Also, consider using
Leonid Keselman's
Jekyll fork
of this page.
|
|